It’s been almost three years since Google’s DeepMind announced a discovery: the AlphaFold neural network has successfully determined the three-dimensional structure of almost all proteins known to date. The discovery was not just a breakthrough in the field of protein folding. It has also demonstrated the potential of machine learning and artificial intelligence in medicine.
This same knowledge made itself felt in a recent study that Published in the journal Nature It was taken advantage of by researchers at the University of Washington. Instead of calculating the structure of existing proteins from sequences of bases, she had an artificial intelligence create new proteins. They are enzymes called luciferases: these are the enzymes that cause bioluminescence through a chemical reaction. Among other things, they make fireflies glow.
“We’ve been able to develop highly efficient enzymes from scratch on a computer rather than relying on enzymes found in nature. This breakthrough means, in principle, that customized enzymes can be developed for almost any chemical reaction,” says molecular biologist Andy Hsien. Wei Yeh, who led the study.
The AI detects 1,600 identical protein structures
The new enzymes could be used, among other things, in diagnostic imaging – for example to examine processes in cells or in medicine to show biochemical reactions that are difficult or impossible to do with conventional methods. Diagnostics using bioluminescence is a relatively new technology, but is said to have great potential. In other areas too, for example in recycling, people are looking for new enzymes that can perform specific tasks.
Finding new enzymes from scratch has always been challenging. In order to become vigorous, they need not only the right nutrient medium, the right substrate, but also ideal conditions in terms of temperature and chemicals. Above all, enzymes only function at will if they are able to dock with the correct proteins, which in turn depends on the three-dimensional structure of the proteins.
In their study, the researchers used diphenylterazine (DTZ), a substance used in organisms to generate light, as a target substrate. The next step was to find enzymes that could dock into DTZs in order to catalyze the chemical reaction that causes bioluminescence. This is where artificial intelligence began: using trRosetta, a neural network designed to predict protein structures, the researchers were able to “hallucinctio” about 1,600 protein scaffolds that could interact with DTZ.
Luminous cells can be seen with the naked eye
To test the results, the team carefully encoded the proposed scaffolds in DNA and then injected them into bacteria to see if the enzymes proposed by the AI actually produced light when combined with DTZ.
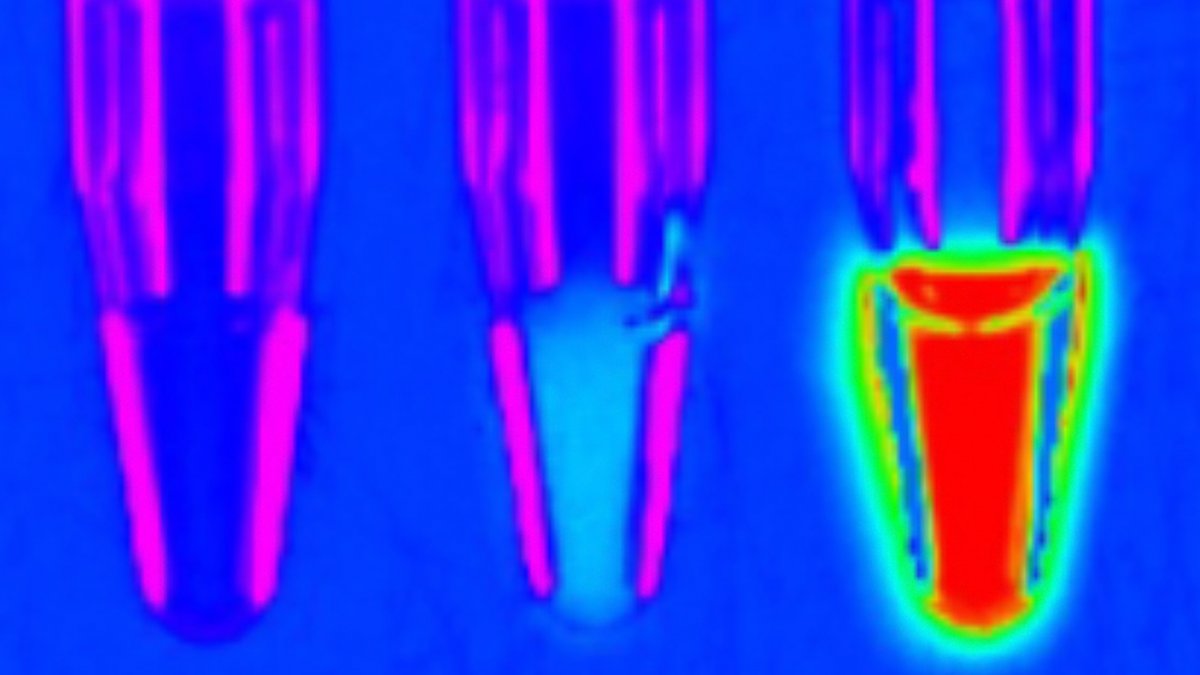
In the end, the “winner” was the new luciferase, which experts dubbed LuxSit (Latin for “Let there be light”). Another improved version later, LuxSit-i, made the cells glow so intensely that the light could be seen with the naked eye. It was also very stable and retained its structure even at 95°C.
“Developing high-potency, specific biocatalysts from scratch with broad applications in biomedicine is an important milestone in computational enzyme design,” says the study, published in Nature, which has already undergone a peer-review process. With the help of neural networks, it is possible to develop specific enzymes that can perform a variety of different tasks and make cell processes visible by emitting different types of light.

(Bachelor’s)

“Total coffee aficionado. Travel buff. Music ninja. Bacon nerd. Beeraholic.”










More Stories
Artemis 2’s Orion Heat Shield Faces Critical Test as Astronauts Prepare for Fiery Earth Reentry
Coral Seeding: Artificial Insemination Makes Coral More Heat Tolerant
Fear, Anger, and Denial: How People Respond to Climate Change – Research